METHODS
A regression framework for brain
network distance metrics
Chal E. Tomlinson1
, Paul J. Laurienti2,3, Robert G. Lyday2,3, and Sean L. Simpson2,4
1Department of Biostatistics, University of North Carolina at Chapel Hill, Chapel Hill, NC, USA
2Laboratory for Complex Brain Networks, Wake Forest School of Medicine, Winston-Salem, NC, USA
3Department of Radiology, Wake Forest School of Medicine, Winston-Salem, NC, USA
4Department of Biostatistics and Data Science, Wake Forest School of Medicine, Winston-Salem, NC, USA
Keywords: Graph theory, Connectivity, fMRI, Small-world, Neuroimaging, Jaccard, Kolmogorov–
Smirnov
a n o p e n a c c e s s
j o u r n a l
ABSTRACT
Analyzing brain networks has long been a prominent research topic in neuroimaging.
However, statistical methods to detect differences between these networks and relate them to
phenotypic traits are still sorely needed. Our previous work developed a novel permutation
testing framework to detect differences between two groups. Here we advance that work to
allow both assessing differences by continuous phenotypes and controlling for confounding
variables. To achieve this, we propose an innovative regression framework to relate distances
(or similarities) between brain network features to functions of absolute differences in
continuous covariates and indicators of difference for categorical variables. We explore
several similarity metrics for comparing distances (or similarities) between connection
matrices, and adapt several standard methods for estimation and inference within our
framework: standard F test, F test with individual level effects (ILE), feasible generalized least
squares (FGLS), and permutation. Via simulation studies, we assess all approaches for
estimation and inference while comparing them with existing multivariate distance matrix
regression (MDMR) methods. We then illustrate the utility of our framework by analyzing the
relationship between fluid intelligence and brain network distances in Human Connectome
Project (HCP) data.
Citation: Tomlinson, C. E., Laurienti,
P. J., Lyday, R. G., & Simpson, S. L.
(2022). A regression framework for
brain network distance metrics.
Network Neuroscience, 6(1), 49–68.
https://doi.org/10.1162/netn_a_00214
DOI:
https://doi.org/10.1162/netn_a_00214
Supporting Information:
https://doi.org/10.1162/netn_a_00214
Received: 18 June 2021
Accepted: 25 October 2021
Competing Interests: The authors have
declared that no competing interests
exist.
INTRODUCTION
Corresponding Author:
Sean L. Simpson
slsimpso@wakehealth.edu
Handling Editor:
Bratislav Misic
Copyright: © 2021
Massachusetts Institute of Technology
Published under a Creative Commons
Attribution 4.0 International
(CC BY 4.0) license
The MIT Press
As brain network analyses have become popular in recent years, neuroimaging researchers
often face the need to statistically compare brain networks (Simpson et al., 2013a). Many
approaches for relating brain networks to clinical outcomes or demographical variables have
been developed. Such methods include, but are not limited to, traditional network models
(e.g., exponential random graph models; Lehmann et al., 2021; Simpson et al., 2011,
2012), tensor regression works on brain network (e.g., Zhang et al., 2018, 2019), Bayesian
approaches (e.g., Dai & Guo, 2017; Wang et al., 2017), statistical learning techniques
(Craddock et al., 2015; Varoquaux & Craddock, 2013; Xia et al., 2020), and testing based
on distance correlation (Székely et al., 2007; Székely & Rizzo, 2009). Despite the advances
made, analysis methods are still needed that enable the comparison of networks while incor-
porating topological features inherent in each individual’s network. To develop such an
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
analysis, we can exploit the fact that brain networks often exhibit consistent organizations
across subjects. For example, a number of studies have reported that nodes with particular
characteristics (e.g., high degree) tend to coincide at the same spatial locations across subjects
(Hagmann et al., 2008; Hayasaka & Laurienti, 2010; Moussa et al., 2011; van den Heuvel
et al., 2008). Although the set of such nodes may not be exactly the same across subjects, there
are large areas of overlap (Hagmann et al., 2008; Hayasaka & Laurienti, 2010). Furthermore,
our study on network modules, or communities of highly interconnected nodes, indicated
that some building blocks of resting-state functional brain networks exhibited remarkable
consistency across subjects (Moussa et al., 2012). It has also been shown that such consistent
organizations differ under different cognitive states (Deuker et al., 2009; Moussa et al., 2011;
Rzucidlo et al., 2013) or in different groups of subjects (Bassett & Bullmore, 2009; Burdette
et al., 2010; Liu et al., 2008; Meunier et al., 2009a; Rombouts et al., 2005; Stam et al., 2007;
Yuan et al., 2010). Thus, an analysis method sensitive to such differences in spatial locations
or patterns can assess network differences across the entire brain (as opposed to univariate
edge-by-edge or node-by-node comparisons). Toward this end, in previous work we
developed a permutation testing framework that detects whether the spatial location of
network features (such as the location of high degree nodes) mapped back into brain space
differs between two groups of networks, and whether distributions of topological properties
vary by group (Simpson et al., 2013b). Despite the utility of this method, it has two major
weaknesses. First, it cannot account for confounding variables. This means that while we
can compare maps of hub regions, for example, between two populations, we cannot control
for differences in characteristics such as age or education. Second, the method relies on
dichotomous grouping to make comparisons. When making comparisons between groups with
clear divisions (male vs. female), this is not an issue. However, it is impossible to assess if there is
a significant relationship between network hub location and continuous measures, such as
intelligence quotient (IQ) score or age.
To address these limitations, we propose an innovative regression framework to relate dis-
tances between brain network features to functions of absolute differences in continuous
covariates and indicators of difference for categorical variables. We will consider several
different types of metrics for establishing distances (i.e., similarity/dissimilarity) between
networks. The first type compares degree distributions. We accomplish this by summarizing
similarities in nodal cumulative degree distributions across multiple networks with the
Kolmogorov–Smirnov statistic (K-S statistic), a measure that quantifies the distance between
two cumulative distribution functions (Kolmogorov, 1933; Smirnov, 1948). The second type
takes into account consistency of key node sets. We do so by summarizing similarities in node
sets across multiple networks with the Jaccard distance (or Jaccard index), a metric that quan-
tifies difference (or similarity) in partitions of a set (Joyce et al., 2010; Meunier et al., 2009b).
The third type of metric measures similarity of nodal degree by employing Minkowski or
Canberra distances between nodal degree vectors (Lance & Williams, 1966).
Within our regression framework we adapt several methods for estimation and inference:
standard F test, F test with individual level effects (ILE), feasible generalized least squares
(FGLS), and permutation. Each observation in the regression framework includes a “distance”
between two individuals, so observations that share individuals are correlated. Thus, the stan-
dard F test is generally not appropriate but presented for comparison. The other methods
attempt to deal with this correlation: (a) F test with ILE—including fixed individual level effects
within the regression, possibly rendering the F test valid; (b) mixed model—including random
individual level effects; (c) FGLS—proposing an artful way to estimate the covariance matrix
(Aitken, 1936); and (d) Permutation—similarity and distance metrics have unknown
50
Permutation testing framework:
A method to test statistical
significance by permuting labels.
Mixed model:
Statistical model containing both
fixed (population-level) and random
(individual-level) effects used to
model multivariate data.
Network Neuroscience
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
Degree distribution:
The probability distribution of the
degree of nodes across a network
(np
2 ): mathematical notation for
“np choose 2,” which represents the
number of pairs one could choose
amongst the np, number of
participants.
distributions; thus, a permutation test may be most appropriate (Simpson et al., 2013b). Per-
mutation tests for these models require permuting the residuals, and we will adapt recent
methods to implement this approach (Kherad-Pajouh & Renaud, 2010, 2014).
In this paper, we detail our proposed regression framework, and discuss several methods for
estimation and inference to be used on a variety of network similarity/dissimilarity metrics. We
assess all combinations of methods and metrics within this framework by using simulated fMRI
data with known differences in connectivity matrix distributions. We then apply our frame-
work to functional brain networks derived from the HCP dataset to investigate the relationship
between fluid intelligence and network distances after accounting for known confounders.
METHODS
Please note the following notational choices: bold font is used to denote vectors or matrices, n =
(np
2 ) = number of observations, np = number of participants, nn = number of nodes, p = number
of covariates (including intercept, if included).
Step 1: Network Construction
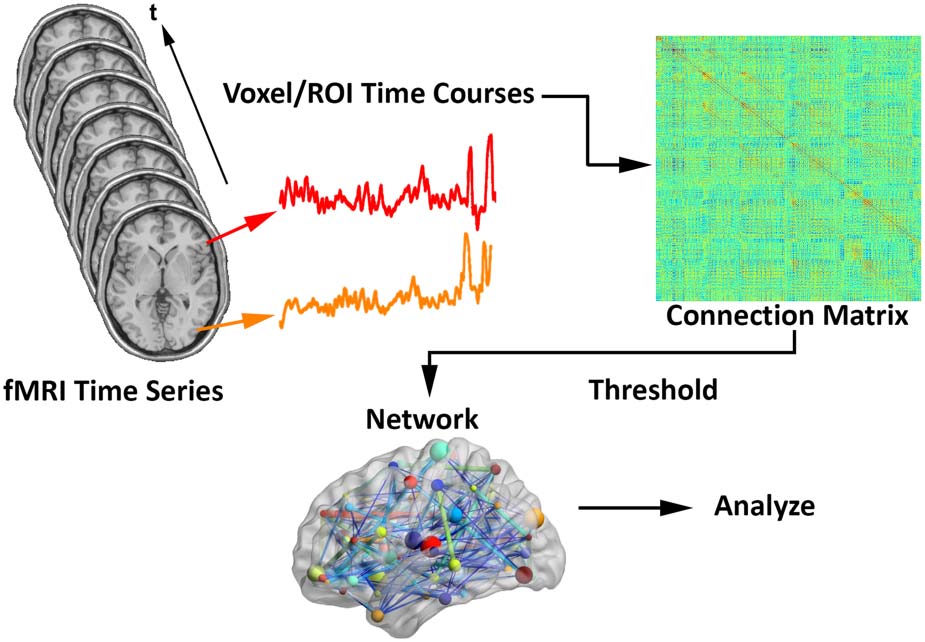
Assuming fMRI connection matrices have already been obtained (see Figure 1), let Ci represent
a weighted nn × nn connection matrix for individual i, with matrix entries ranging from 0 (indi-
cating no connection between the respective nodes) to 1 (strongest connection). We only con-
sidered undirected networks, so matrices were symmetric, with the number of row = number of
columns = number of nodes (methods are easily adaptable if directed networks are desired).
Let bi represent an nn × 1 binary key node vector for individual i, with an entry of 1 repre-
senting a key node and 0 a node that is not key. As stated in our previous work (Simpson et al.,
2013b), key nodes can be identified based on nodal characteristics such as high degree, high
centrality, or other desired characteristics. Since key nodes were compared across subjects, it
was important to employ the same criterion in all of the networks (e.g., top 10% highest cen-
trality, node degree >200, etc.). Alternatively, key nodes can be identified as those belonging
to a particular module or network community, a collection of highly interconnected nodes.
The resulting key nodes would form a set, and the consistency of the spatial location of the
Schematic for generating brain networks from fMRI time series data—recreated from
Figure 1.
(Fornito et al., 2012; Simpson et al., 2013a). Functional connectivity between brain areas is esti-
mated based on time series pairs to produce a connection matrix. A threshold is commonly applied
to the matrix to remove negative and/or “weak” connections.
Network Neuroscience
51
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
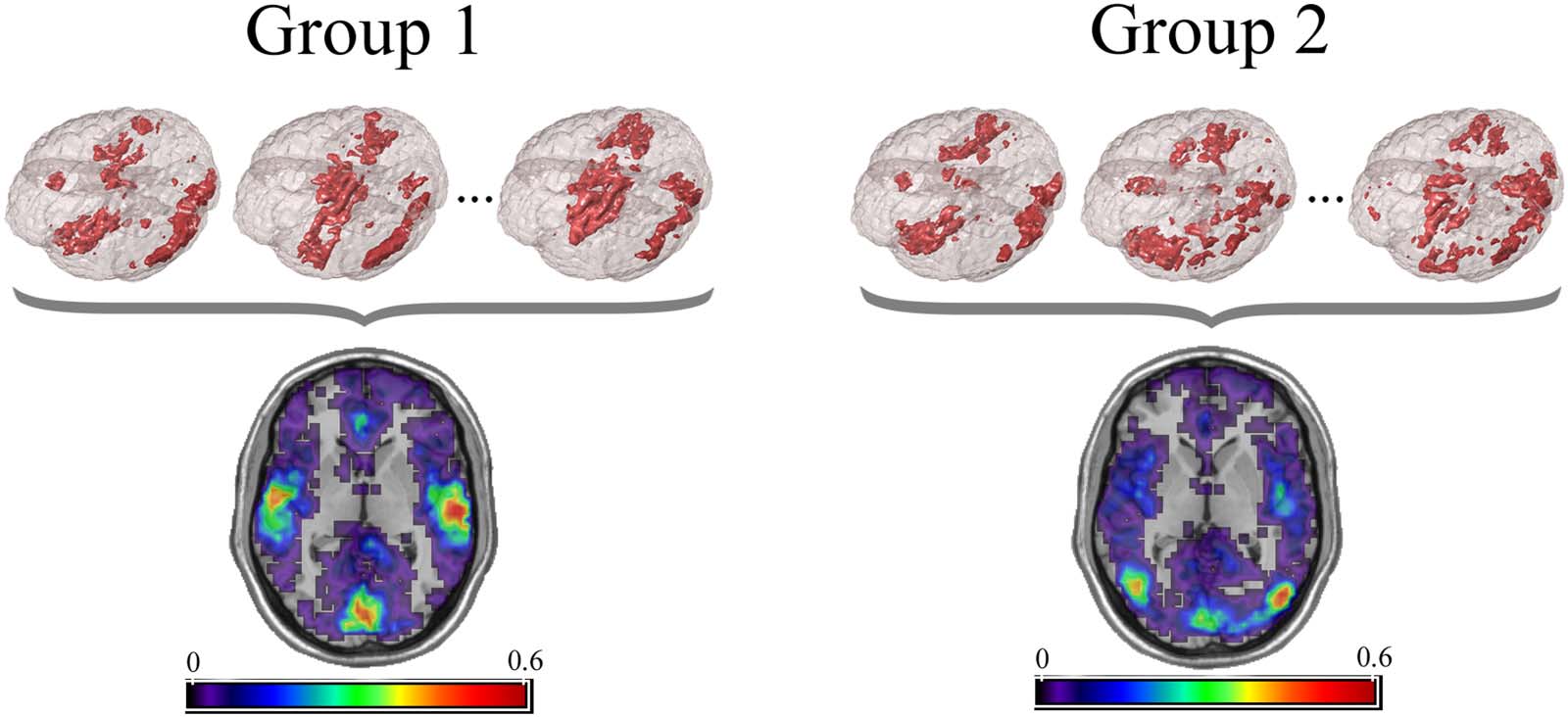
Example visualization of key node sets from voxel-based networks in brain space. The 3D brain images (top) are three represen-
Figure 2.
tative subjects from each group with the key node sets defined to be those with the top 20% highest degree. Overlap maps (bottom) show the
proportion of subjects with key nodes in given areas. Figure recreated from Simpson et al. (2013b).
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
nodes could be compared across groups. An example visualization of key node sets from
voxel-based networks in brain space is given in Figure 2.
We constructed an nn × 1 weighted nodal degree vector di by summing Ci over its rows (or
equivalently, columns). Each entry in vector di represented the “degree” of its respective node
for individual i. It should be noted here that when the binary key node vector has to do with
node degree, say 20% highest degree as will used later, bi is just a binarized version of di.
As was described for the binary key node vector in the last paragraph, other weighted nodal
vectors could have been used. For simplicity, we focused on degree within the methods and
simulation sections. It should also be noted that none of the methods employed here are spe-
cific to nodal vectors. That is, these methods could also be implemented on differences
between connection matrices, for example.
Step 2: Establish Similarity/Dissimilarity Between Networks
This section covers some of the metrics we used to gauge distances between individual net-
works given the insight they can provide into brain network organizational differences.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
Kolmogorov–Smirnov (KS) statistic. Degree distributions, which help quantify the topology of
networks, are likely more similar within distinctive groups than they are between these groups.
We employed the log of the KS statistic to quantify this potential dissimilarity.
(cid:1)
log KSij
(cid:3)
(cid:1)
(cid:4)
(cid:4)
(cid:4)
(cid:4)
¼ log supx Fi xð Þ − Fj xð Þ
(cid:3)
Empirical distribution function:
Estimate of the cumulative
distribution function using the data.
KSij, a scalar, is the Kolmogorov–Smirnov statistic between nodal degree distributions for indi-
viduals i and j. Fi(x) represents the empirical distribution function for observations from the
nodal degree vector di. So, supx|Fi(x) − Fj(x)| gives the biggest difference between the
empirical degree distributions between individuals i and j. Bigger values indicate more
dissimilarity.
Logarithmic transformation:
Taking the log of a variable.
A note on the logarithmic transformation of the KS statistic: when all distances are nonneg-
ative, it is common practice to take a log transformation. Within our simulations, KS was the
Network Neuroscience
52
A regression framework for brain network distance metrics
Power:
Probability of correctly rejecting the
null hypothesis.
only metric that saw improvements in power or type I error when taking such a transformation.
For ease of interpretability, none of the other distances presented here utilized a logarithmic
transformation.
JDij ¼ M01 þ M10
M11 þ M01 þ M10
JDij, a scalar, is the Jaccard distance ( JD) between two (binary) key node
Jaccard distance.
vectors of individuals i and j, bi, and bj. M11 is the number of nodes such that bi and bj both
have a value of 1, M01 is the number of nodes where bi = 0 and bj = 1, and M10 is the number
of nodes where bi = 1 and bj = 0. JDij gives the proportion of key nodes (in either set) that do
not share key status between individuals i and j. Values of JDij range from 0 (perfect overlap)
to 1 (no overlap).
JIij ¼ 1 − JDij ¼
M11
M11 þ M01 þ M10
JIij, a scalar, is the Jaccard index ( JI) between two (binary) key node vectors of
Jaccard index.
individuals i and j, bi, and bj. JIij gives the proportion of nodes that share key status between
individuals i and j. Values of JIij range from 1 (perfect overlap—key node networks that are the
same) to 0 (no overlap). Although the JI simply equals 1 − JDij, it is included here to compare a
distance and similarity metric.
(cid:5)
(cid:5)
Eij ¼ di − dj
(cid:5)
(cid:5)
2
¼
X# of nodes
k¼1
!1
2
jdi k½
(cid:2) − dj k½
(cid:2)j2
Euclidean distance. Eij, a scalar, is the Euclidean distance between nodal degree vector i and
nodal degree vector j, where di[k] represents the degree of node k for individual i. Bigger
values of Euclidean distance indicate more dissimilarity.
See the Supporting Information for Minkowski distance and Canberra distance.
Step 3: Evaluating Differences Between Networks
Distij ¼ X T
ij;conβcon þ X T
ij;coi
β
coi
þ εij
Standard F test. Distij represents the distance between nodal vectors of individuals i and j.
Distij is a generic placeholder for any metric outlined previously in Step 2, that is, JD ( JDij), KS
statistic (KSij), Minkowski distance (Mp
ij ), or Canberra distance (Cij).
X T
ij;con, an 1 × (p − 1) vector, contains the intercept and differences in confounding covar-
iates (e.g., for our data, sex, educational attainment, age, and body mass index) between indi-
viduals i and j (with corresponding unknown ( p − 1) × 1 parameter vector β
con) to control for
differences that may confound the relationship between the covariate of interest and the given
distance.
X T
ij;coi, a scalar, contains the difference in the covariate of interest (or an indicator of different
group membership for group-based analyses) between individuals i and j (with corresponding
unknown parameter βcoi).
Network Neuroscience
53
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
Splitting the design matrix X, an n × p matrix, into confounding and of interest covariates is
ij;coi can be combined into the 1 × p vector X T
purely a notational preference. X T
(with corresponding unknown p × 1 parameter vector β).
ij;con and X T
ij
εij accounts for the random error in the distance ( Jaccard, KS, etc.) value. If εij were inde-
pendent, homoscedastic, and approximately normally distributed, the F test of a standard lin-
ear regression would be an appropriate test. We expect correlation among observations from
the same individual, so we included this standard testing procedure here mainly for
comparison.
As an example, to test whether there is an association between IQ (continuous) and the
spatial consistency of hub nodes (top 20% highest degree) after controlling for age (continu-
ous), sex (binary), and treatment (binary) status, our model would be
(cid:7)
(cid:6)
þ 1 Sexi
(cid:6)
þ 1 Trti
6¼ Sexj
6¼ Trtj
þ IQi − IQj
þ Agei − Agej
JDij ¼ β
þ εij
(cid:4)
(cid:4)
(cid:4)β
(cid:4)
(cid:4)β
(cid:4)
(cid:4)
(cid:4)
β
(cid:7)
(cid:4)
(cid:4)
β
3
0
1
2
4
with the associated null hypothesis H0: β4 = 0.
Dist ¼ X T β þ ID1α1 þ … þ IDnp
αnp
þ (cid:2)
Standard F test with individual level fixed effects (F test with ILE). Dist is an n × 1 vector of known
distance metrics (as outlined in Step 2), XT is the n × p design matrix (intercept optional) of
known covariates with corresponding p × 1 unknown parameter vector β, IDi is the n × 1
known indicator variable for individual i with corresponding unknown parameter αi. By
accounting for individual level effects, we hope to induce independence, homoscedasticity,
and normality for the error terms within the n × 1 random vector (cid:2). This would (potentially)
allow for an F test to appropriately evaluate the covariates of interest.
Random effect:
Variable whose effect varies across
individuals.
Fixed effects:
Variables whose effects are constant
across individuals.
It should be mentioned that we attempted a mixed model formulation
Mixed model approach.
where each individual had their own random effect and all other covariates had fixed effects
(where Dist = XTβ + ID1b1 + ⋯ + IDnpbnp + (cid:2) and bi ∼ N(0, g) for all i). We also attempted bi ∼
N(0, gi) for all i. Unfortunately, these calculations were too computationally intensive in the
simulations we ran (using the lmer function of the R package lme4; Bates et al., 2015). Instead,
we tried a generalized least squares approach outlined in the next section.
Dist ¼ X T β þ (cid:2)
Feasible generalized least squares. Dist and XTβ (intercept included) are as before, but instead
we assume (cid:2) ∼ (0, (cid:2)), where (cid:2) is the n × n covariance matrix. Generalized least squares (GLS)
allows for estimating parameters when there is correlation among the residuals in ordinary
least squares regression. However, GLS requires (cid:2) to be known. An unrestricted n × n covari-
ance matrix has n(n + 1)/2 parameters to estimate. This is infeasible as we only have n obser-
vations. Thus, we restricted the form of (cid:2) in order to estimate it. For a detailed explanation,
please see Supporting Information.
Permutation test. A permutation test requires no knowledge of how the test statistic of interest
is distributed under the null hypothesis (e.g., H0: no significant difference among IQ). The dis-
tribution under the null hypothesis is empirically “generated” by permuting data labels. We
employed the Freedman–Lane approach (Freedman & Lane, 1983; Kherad-Pajouh & Renaud,
2010) using the lmperm function in the R package permuco (Frossard & Renaud, 2019a) with
10,000 permutations, while permuting across individuals to preserve exchangeability. For a
Network Neuroscience
54
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
detailed explanation, please see Supporting Information as well as the package vignette for
permuco (Frossard & Renaud, 2019b).
Multivariate distance matrix regression. Multivariate distance matrix regression (MDMR) is an
existing method that has been included here for comparison. It tests the significance of asso-
ciations of response profile (dis)similarities and a set of predictors. Originally this was done
using only permutation (Anderson, 2001), but has been extended to analytic p values as well
(McArtor et al., 2017). MDMR was run using the mdmr function in the MDMR package in R
(McArtor, 2018). Both the permutation and analytic versions were run with the np × np dis-
tance matrix D (the distance matrix analog of Dist) and the np × p design matrix Xp (covariates
of interest for each participant). The permutation method was run with 10,000 permutations.
SIMULATION STUDIES
The following simulation study is done using a factorial approach. There are four different set-
tings (named Simulations 1–4). For each simulation setting, we explore four different metrics
(KS, JD, JI, and Euclidean). For each simulation and metric combination, six different methods
are considered (permutation, F test, FGLS, F test ILE, MDMR analytic, and MDMR permuta-
tion). Subsequent sections will present details and results for each of these “factors.”
Data
We conducted four simulation settings to assess how well our proposed approaches could
detect various relationships between brain network properties and covariates of interest. Each
simulation contained 100 subjects, with four covariates of interest. A fair coin was flipped
for each subject to determine their sex (SEX = male or female). Half of subjects were
assigned to treatment and the other half were assigned to placebo (TRT = treatment or pla-
cebo). IQ and age were both simulated from a normal distribution with mean of 100 and a
standard deviation of 15 (rounded to the nearest integer). This resulted in two binary (SEX,
TRT ), and two continuous (AGE, IQ) covariates—variables were given names purely for pur-
poses of explication.
For Simulations 1–3, we simulated fMRI connectivity matrices with 268 nodes each to
mimic the experimental data detailed in the next section. In each simulation, a 268 × 268
symmetric matrix (with entries ranging from 0 to 1) was generated for each subject. Entries
within each connectivity matrix were drawn from three types of distributions: (a) a low-
connectivity noise beta distribution, Beta(4
3, 6); (b) a high-connectivity noise distribution,
Beta(4, 6); and (c) a signal distribution dependent on covariates and signal percentage, Beta(sp ·
ai + (1 − sp) · 4
3, 4 + (IQ1 − 100) * .2 + (Trti = = “Treatment”) * 6) represented the
covariate-dependent parameter and sp represented the signal percentage (from 0 to 100%).
When the signal percent (sp) was 100%, Beta(sp · ai + (1 − sp) · 4
3, 6) = Beta(ai, 6). Similarly, when
signal percent was 0%, Beta(sp · ai + (1 − sp) · 4
3, 6) = Beta(4
3, 6), and was therefore identical to the
noise distribution and no longer dependent on covariates.
3, 6), where ai = max(4
Simulation 1 had a 15 × 15 region where all individuals had the same high-connectivity
noise distribution, a 15 × 15 region where the signal distribution was dependent on covariates,
and the rest of the 268 × 268 connection matrix was drawn from the low-connectivity noise
distribution. Simulation 2 was the same as 1, but the signal distribution dependent on covar-
iates was moved to expand the border of the 15 × 15 high-connectivity signal distribution to a
combined 21 × 21 region. Simulation 3 was the same as 1, with two additional 15 × 15 regions
where all individuals had the same high-connectivity distribution. For a drawn to scale
Fair coin:
A coin which has a 50% chance of
landing on either side.
Symmetric matrix:
A matrix which entry aij = aji for all
i and j. In the connectivity matrix
space, this lets us know we are
considering an undirected network.
Beta distribution:
A family of continuous probability
distributions defined on the closed
interval [0, 1].
Network Neuroscience
55
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
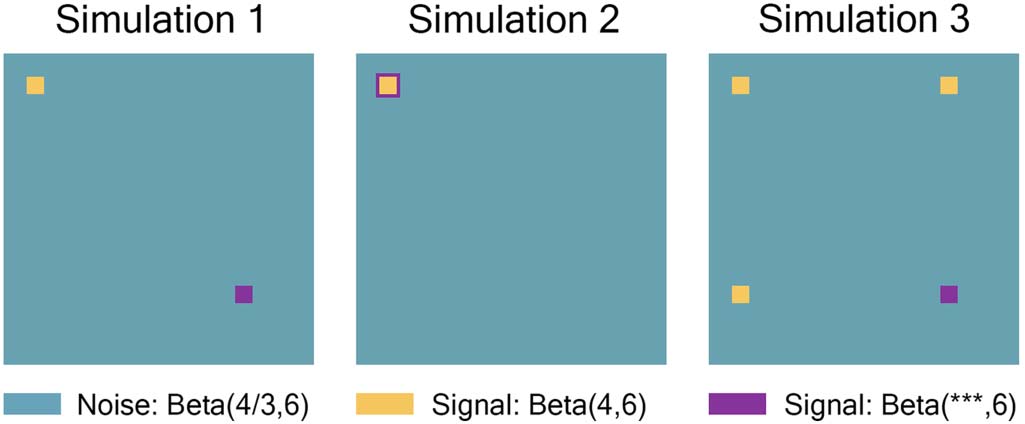
Figure 3. Simulation 1 had a 15 × 15 region where all individuals had the same high-connectivity
noise distribution (shown in yellow), a 15 × 15 region where the signal distribution was dependent
on covariates (shown in purple), and the rest of the 268 × 268 connection matrix was drawn from
the low-connectivity noise distribution (shown in teal). Simulation 2 was the same as 1, but the
signal distribution dependent on covariates was moved to expand the border of the 15 × 15
high-connectivity signal distribution to a combined 21 × 21 region. Simulation 3 was the same
as 1, with two additional 15 × 15 regions where all individuals had the same high-connectivity
distribution. It should be noted that we assumed the connectivity matrices to be symmetric, with
0 entries along the diagonal. The plots were drawn to scale.
representation of these simulations, see Figure 3. It should be noted that we assumed the con-
nectivity matrices to be symmetric, with 0 entries along the diagonal.
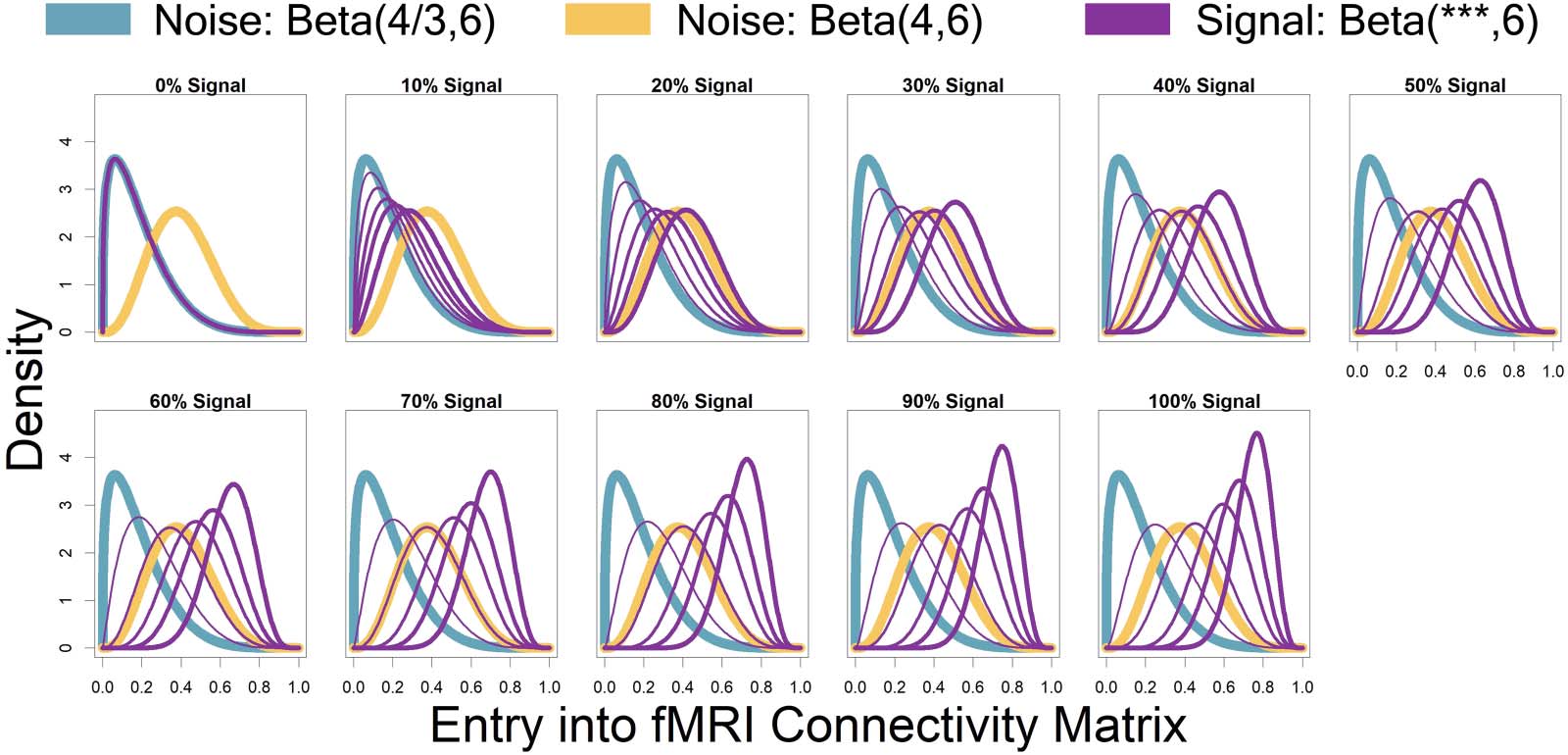
Represented by the same colors from Figure 3 simulated connectivity matrices, Figure 4
displays the distributions used for those matrices as signal percentage increased. The low-
connectivity noise distribution was distributed Beta(4/3, 6) and was shown in teal (not affected
by signal percentage). The high-connectivity noise distribution was distributed Beta(4, 6) and is
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
/
t
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
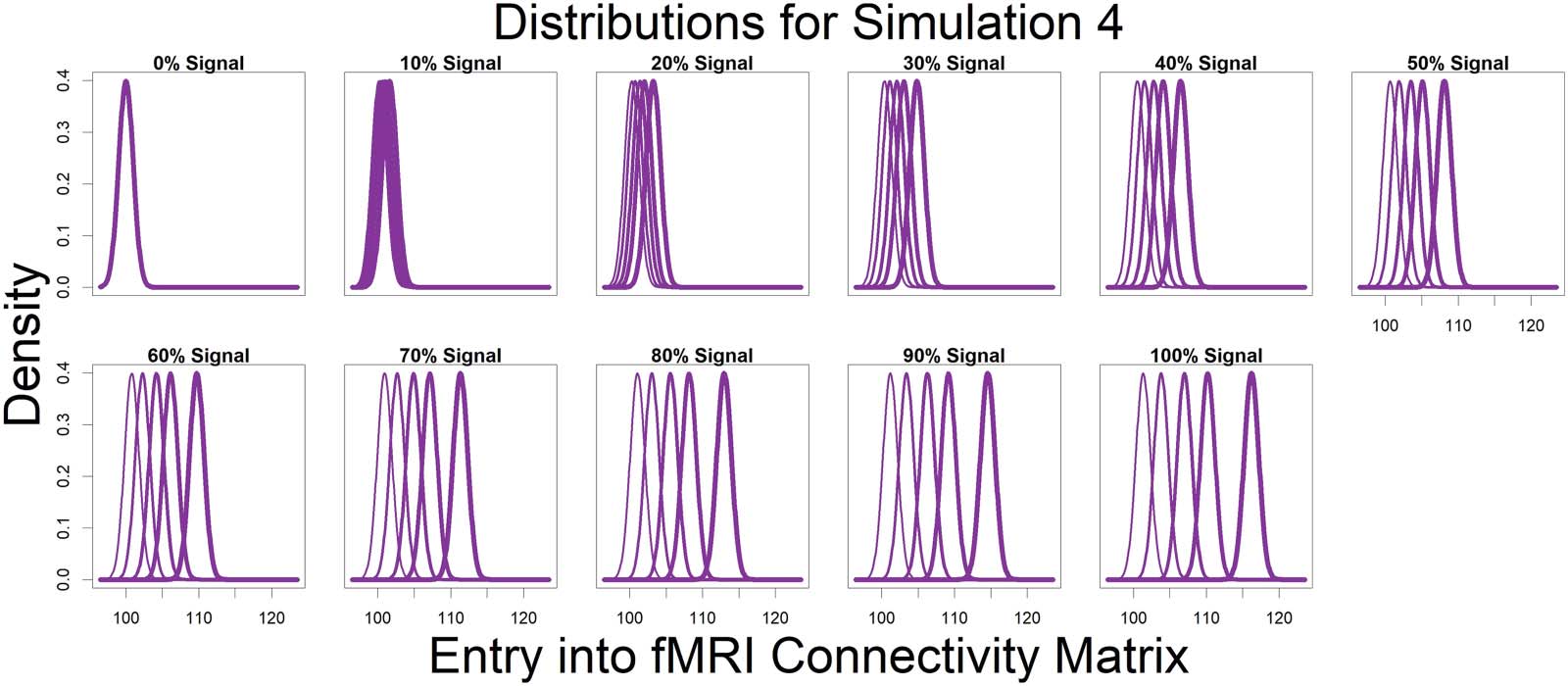
Figure 4. Represented by the same colors from Figure 3 simulated connectivity matrices, displayed here are the distributions used for those
matrices as signal percentage increased. The low-connectivity noise distribution was distributed Beta(4/3, 6) and was shown in teal (not
affected by signal percentage). The high-connectivity noise distribution was distributed Beta(4, 6) and is in yellow (not affected by signal
percentage). The “covariate-dependent” signal region can be seen in purple and was distributed Beta(***, 6). The *** parameter had some
distribution based on the underlying covariate distribution and signal percentage. There are five purple distributions in each plot representing
the 0.01, 0.25, 0.5, 0.75, and 0.99 quantiles (shown in increasing thickness) from the *** distribution (for example, the 0.25 quantile distri-
bution is represented by an individual with an IQ of 69 and a “Treatment” status or an individual with an IQ of 99 and a “Placebo” status; the
0.75 quantile distribution is represented by an individual with an IQ of 101 and a “Treatment” status or an individual with an IQ of 131 and a
“Placebo” status). Furthermore, the “covariate-dependent” (purple) signal region’s distribution goes from being the same as the low-
connectivity noise region’s distribution (at 0% signal) to more and more different than the noise region’s distribution as signal percentage
increases.
Network Neuroscience
56
A regression framework for brain network distance metrics
in yellow (not affected by signal percentage). The “covariate-dependent” signal region can be
seen in purple and was distributed Beta(***, 6). The *** parameter had some distribution based
on the underlying covariate distribution and signal percentage. There are five purple distribu-
tions in each plot representing the 0.01, 0.25, 0.5, 0.75, and 0.99 quantiles (shown in increasing
thickness) from the *** distribution (for example, the 0.25 quantile distribution is represented by
an individual with an IQ of 69 and a “Treatment” status or an individual with an IQ of 99 and a
“Placebo” status; the 0.75 quantile distribution is represented by an individual with an IQ of 101
and a “Treatment” status or an individual with an IQ of 131 and a “Placebo” status). Furthermore,
the “covariate-dependent” (purple) signal region’s distribution goes from being the same as the
low-connectivity noise region’s distribution (at 0% signal) to more and more different than the
noise region’s distribution as signal percentage increases.
For Simulation 4, we simulated nodal degree vectors of length 268 (instead of 268 × 268
connectivity matrices as in Simulations 1–3) to assess the method’s ability to detect distribu-
tional differences rather than location differences. All entries of each individual’s degree vector
were simulated Normal(100 + sp · ai, 1), where sp (signal percent) and ai (covariate-dependent
parameter for individual i) were as described with Simulations 1–3. Within Figure 5, there were
five purple distributions in each plot, representing the 0.01, 0.25, 0.5, 0.75, and 0.99 quantiles
(shown in increasing thickness) from the distribution of the mean parameter, 100 + sp · ai. At
0% signal, all individual’s nodal degree vectors were drawn from Normal(100, 1). As signal
percentage increased, the mean parameter of the covariate-dependent distributions became
more and more distinct.
Results
We assessed methods with the four simulation scenarios detailed in the previous section. Each
simulation was run 10,000 times. Nodal degree vectors (used for the KS and Euclidean dis-
tances) were created by summing across rows of the connectivity matrices. Key nodes of inter-
est (binary degree vectors used for the JD and JI) based on node degree were identified,
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
Figure 5. For simulation 4, we simulated nodal degree vectors of length 268 (instead of 268 × 268 connectivity matrices as in Simulations 1–3)
to assess the method’s ability to detect distributional differences rather than location differences. All entries of each individual’s degree vector
were simulated Normal(100 + sp · ai, 1), where sp (signal percent) and ai (covariate-dependent parameter for individual i) were as described
with Simulations 1–3. There were five purple distributions in each plot, representing the 0.01, 0.25, 0.5, 0.75, and 0.99 quantiles (shown in
increasing thickness) from the distribution of the mean parameter, 100 + sp · ai. At 0% signal, all individual’s nodal degree vectors were
drawn from Normal(100, 1). As signal percentage increased, the mean parameter of the covariate-dependent distributions became more and
more distinct.
Network Neuroscience
57
A regression framework for brain network distance metrics
selecting the top 20% highest degree (hub) nodes and mapping those to 1 while mapping all
remaining nodes to 0. The KS statistic and Euclidean distance were calculated for each pair of
individuals using their nodal degree vectors. The JD and JI were calculated for each pair of
individuals using their binary degree vectors.
The percentages of p values less than α = 0.05 for the covariates of interest were recorded
for each combination of signal percent (0%, 10%, …, 100%), distance metric (KS, JD, JI, Euclid-
ean), and testing framework (F test, permutation, GLS, F test with individual level effects,
MDMR analytic and MDMR permutation). In this section, we discuss whether type I error
rate was controlled and at what signal percent the 80% power threshold was reached. Then,
we compare results with MDMR. For a visual display of the results, see Figure 6. For more plots
and a visual display with additional distance metrics, please see Supporting Information. The
standard F test did not control type I error when testing age. It was included in the Figure for
reference, but is not mentioned any further in this section.
Type I error rate:
Probability of incorrectly rejecting
the null hypothesis.
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
t
/
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
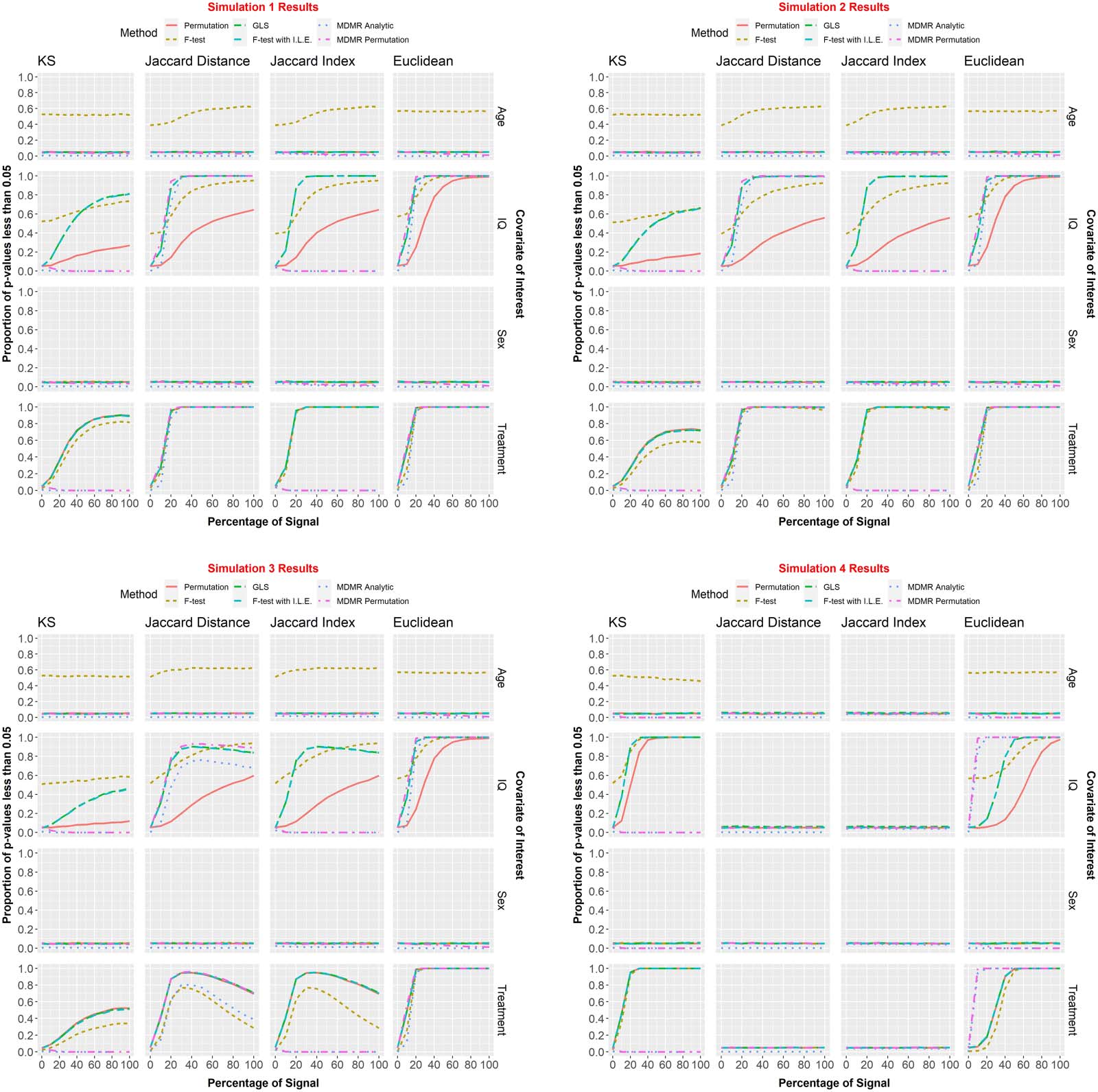
Figure 6. We assessed methods with the four simulation scenarios detailed in the Results section. Each simulation was run 10,000 times. The
percentages of p values less than α = 0.05 for the covariates of interest were recorded for each combination of signal percent (0%, 10%, …,
100%), distance metric (KS, JD, JI, Euclidean), and testing framework (F test, permutation, GLS, F test with individual level effects, MDMR
analytic and MDMR permutation). It should be noted here that Age and Sex are “null” covariates (that have no bearing on the data generating
process) and are included to assess type I error control of the methods on both continuous and categorical variables.
Network Neuroscience
58
A regression framework for brain network distance metrics
Kolmogorov–Smirnov statistic. For the KS metric, all methods (not including F test) adequately
controlled type I error. For Simulation 1, the GLS and F test with ILE methods reached 80%
power within 100% of signal for continuous covariates. The Permutation method never
reached 80% power for continuous covariates. For binary covariates, all four methods reached
80% power within 60% of signal. For Simulations 2–3, no methods reached 80% power at any
level of covariate-dependent signal. For Simulation 4, the GLS and F test with ILE methods
reached the power threshold within 20% of signal. Permutation reached the threshold within
30% of signal. All three methods controlled type I error.
Jaccard distance and Jaccard index. Please note, p values for all methods used within our
regression framework were the same whether the JD or the JI were used (this is not true
of the MDMR methods). For the JD and JI, all methods (not including F test) adequately
controlled type I error. For Simulations 1–3, IQ, the GLS and F test with ILE methods
reached the power threshold within 20–30% of signal; the permutation method never
reached the minimum power requirement. For Simulations 1–3, Treatment, the GLS, F test
with ILE, and Permutation methods reached the power threshold within 20% of signal.
Surprisingly, as signal percentage increased, some situations had decreasing power with
some even falling far back below the power threshold. This only occurred with the Jaccard
metrics. For Simulation 4, no methods could detect significance. All methods’ power levels
hovered around α = 0.05, the significance level. This behavior was expected. For a given
individual, all entries of the nodal degree vector are drawn independently and identically
distributed (iid). Then, highest valued entries of the vector are selected as key nodes.
Because all nodes are iid, the distribution set of “key nodes” is uniform on its support. While
the centrality parameter of the distribution varies from person to person based on their
covariates, this location shift is effectively canceled out by the binarization. Thus, a differ-
ence in the mean of the distributions does not lead to a difference in the distribution of the
key node vectors, and so it is unsurprising that the power in this setting appears to be 5%. It
is not that the Jaccard is causing power issues here, rather, it is because the null is actually
true here.
Euclidean. For the Euclidean metric, all methods (not including F test) adequately controlled
type I error. For Simulations 1–3, IQ, the GLS and F test with ILE methods reached the power
threshold within 20% signal. The permutation method reached the minimum power require-
ment within 50% of signal. For Simulations 1–3, Treatment, all methods reached the power
threshold within 20% signal. For Simulation 4, IQ, the GLS and F test with ILE methods
reached the power threshold within 50% signal. The permutation method reached the mini-
mum power requirement within 80% of signal. For Simulation 4, Treatment, all methods
reached the power threshold within 40% signal.
Comparison with MDMR. For KS and JI, neither MDMR permutation nor MDMR analytic had
adequate power for the KS statistic. Our approaches vastly outperformed them for all simula-
tion scenarios. For Simulations 1–3, JD and Euclidean, the MDMR permutation method had
comparable power with the GLS and F test with ILE methods, while MDMR analytic had
slightly less power. For Simulation 4, Jaccard, see explanation in section JD and JI above.
For Simulation 4, Euclidean, both MDMR methods had considerably more power than any
of the methods within our framework. MDMR analytic type I error rate remained close to 0
for all simulations. MDMR permutation started off with the expected type I error rate of approx-
imately 0.05, but as signal increased, type I error rate approached 0 as well.
Network Neuroscience
59
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
EXPERIMENTAL STUDIES
Data
The HCP data released to date include 1,200 individuals (Van Essen et al., 2013). Of those,
1,113 (606 females; 283 minority) have complete MRI images, cognitive testing, and detailed
demographic information. The 397 subjects used are what remained after quality control
assessment of head motion and global signal changes for both scan types, removal of those
with missing data, and random selection of one individual from each family to ensure
between-subject independence. Participants in the HCP completed two resting-state scans
and two working memory scans. The two scans were collected with different phase encoding
(right to left vs. left to right). The resting-state scans were collected back-to-back while partic-
ipants quietly viewed a fixation point. The 2-back task was a block design that interleaved the
2-back condition with a 0-back condition and a rest period. The working memory task utilized
photos and different blocks had different photo types (faces, body parts, houses, and tools).
Participants were alerted prior to each block to indicate the task type. For the 2-back they were
instructed to respond anytime the current stimulus being presented matched the stimulus two
trials back. The HCP performed extensive testing and development to ensure comparable
imaging at the two sites (Van Essen et al., 2012). The blood oxygenation level-dependent
(BOLD)–weighted images were collected using the following parameters: TR = 720 ms, TE =
33.1 ms, voxel size 2 mm3, 72 slices, 1,200 images.
The main covariate of interest for this analysis was fluid intelligence. Other covariates
included in model formulation were age, body mass index (BMI), education, handedness,
income, alcohol abuse, alcohol dependence, ethnicity, race, sex, and smoking status. For a
summarization and explanation of the variables, see Table 1 and Table 2.
Data Processing and Network Generation
The current project used the minimally processed fMRI data provided by the HCP (Glasser
et al., 2013) for resting-state and working memory. Additional preprocessing steps included
motion correction using ICA-AROMA (Pruim et al., 2015), removal of the first 14 volumes from
each scan, and band-pass filtering (0.009–0.08 Hz) to remove physiological noise and low-
frequency drift. It was necessary to account for the block design of the working memory task.
First, the block design was modeled in SPM12, yielding regressors for the 0-back and rest
blocks as well as the cues. Additionally, since each scan was collected twice with different
phase encoding, the scans were concatenated and a scan-specific regressor was added. All
Table 1.
Summarization and explanation of HCP covariates treated as continuous (within the regression framework)
Age
BMI
Education
Fluid intelligence
Handedness
Income
Mean (SD)
28.7 (3.7)
26.5 (5.1)
15.0 (1.7)
17.3 (4.7)
65.1 (43.6)
5.0 (2.1)
Notes
In years
Body mass index
Integer values 11 to 17 (years pf education completed)
Integer valued from 4 to 24
Values range from −100 to 100 by 5 (−100, −95, …, 95, 100)
SSAGA income score – Total household income: <$10,000 = 1, 10K–19,999 = 2, 20K–29,999 = 3, 30K–39,999 = 4, 40K–49,999 = 5, 50K–74,999 = 6, 75K–99,999 = 7, >=100,000 = 8
Network Neuroscience
60
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
Table 2.
Summarization and explanation of HCP covariates treated as categorical (within the regression framework)
Alcohol abuse
Alcohol dependence
Ethnicity
Race
Sex
54 met the DSM4 criteria for alcohol abuse, 343 did not
28 met the DSM4 criteria for alcohol dependence, 369 did not
41 Hispanic/Latino, 351 Not Hispanic/Latino, 5 Unknown or Not Reported
1 Am. Indian/Alaskan Nat., 31 Asian/Nat. Hawaiian/Other Pacific Is., 48 Black
or African Am., 13 More than one, 10 Unknown or Not Reported, 294 White
207 Female, 190 Male
Smoking status
69 reported as still smoking, 328 did not
regressors, including the total gray matter, white matter, and cerebrospinal fluid signals and
realignment parameters were used in a single regression analysis. The residual signal following
regression of the extraneous variables was retained only for periods aligned with the 2-back
blocks. The blocks of data were then concatenated into a single time series containing 274
BOLD images. Resting-state data followed a similar pipeline, but without the task-specific
regressors. The resulting resting-state data contained 2,372 BOLD images. After preprocessing,
the brain was parcellated into 268 regions as defined in the Shen Atlas (Shen et al., 2013), and
the signal from all voxels within each region was averaged for each participant. A functional
network was constructed for each participant by computing the Pearson (full) correlation
between the resultant time series for each region pair. Negative correlation values were set
to zero because the neurobiological interpretation of positive and negative edges are very dif-
ferent (Parente et al., 2017; Schwarz & McGonigle, 2011), and since the distributions of net-
work variables (such as degree) are different for positive and negative edges (Fraiman et al.,
2009; Saberi et al., 2021). Although negative correlations are not regularly used in network
neuroscience, if they are used, positive and negative networks should be generated and
assessed separately (Schwarz & McGonigle, 2011). For the current work we focused on pos-
itive networks, but one could simply perform an additional analysis on negative networks
if so desired.
Results
Nodal degree vectors (used for the KS and Euclidean distances) were created by summing
across rows of the connectivity matrices. Key nodes of interest (binary degree vectors used
for the JD) based on node degree were identified, selecting the top 20% highest degree
(hub) nodes and mapping those to 1 while mapping all remaining nodes to 0. KS statistic
and Euclidean distance were calculated for each pair of individuals using their nodal degree
vectors. The JD was calculated for each pair of individuals using their binary degree vectors.
Distance covariates for each pair of individuals were calculated. A continuous variable’s
distance (age, for instance) was calculated as |Agei − Agej| for the pair of individuals i and j. A
binary or categorical variable’s distance (education, for instance) was calculated as 1{Edui ≠
Eduj} for the pair of individuals i and j.
We evaluated differences between networks using the standard F test with ILE, as it was the
best in our simulations at controlling type I error and providing sufficient power across all dis-
tance metrics (while also being computationally inexpensive). Parameter and standard error
estimates can be found in Supporting Information Tables S1 and S2. Each parameter estimate
represented the average amount the given brain distance metric (KS, Jaccard, etc.) changed
Network Neuroscience
61
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
t
/
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
P values for HCP resting-state and working memory brain scans when modeled with our
Table 3.
given regression framework and tested using the standard F test with fixed individual level effects
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
Note. Parameter estimates and standard errors can be found in Supporting Information.
based on a one-unit difference in the respective covariate, after controlling for other covari-
ates. A complete list of p values for both resting-state and working memory can be seen in
Table 3. Given the high degree of dependence between these results, and the illustrative and
exploratory nature of our analysis, there have been no adjustments for multiple comparisons.
After adjusting for the other confounding variables, the covariate of interest, fluid intelli-
gence, had a statistically significant relationship with all distance metrics (KS, JD, Euclidean)
for resting-state fMRI, and non-KS distance metrics for working memory fMRI. The level of
significance was considerably higher for the resting-state data. Figure 7A shows that the
resting-state spatial distribution of brain hubs are relatively concentrated in the brain regions
making up the default mode network (known to have high degree at rest). However, there is a
higher concentration of hubs localized to the default mode network in the bottom quartile of
fluid intelligence compared to those in the top quartile. Figure 7B shows that hub locations
during the working memory task shift from default mode areas to regions that make up the
central executive attention network (CEN) more so in the top quartile of fluid intelligence.
Note that in those individuals in the bottom quartile of intelligence, the hubs are most
Network Neuroscience
62
A regression framework for brain network distance metrics
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
t
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
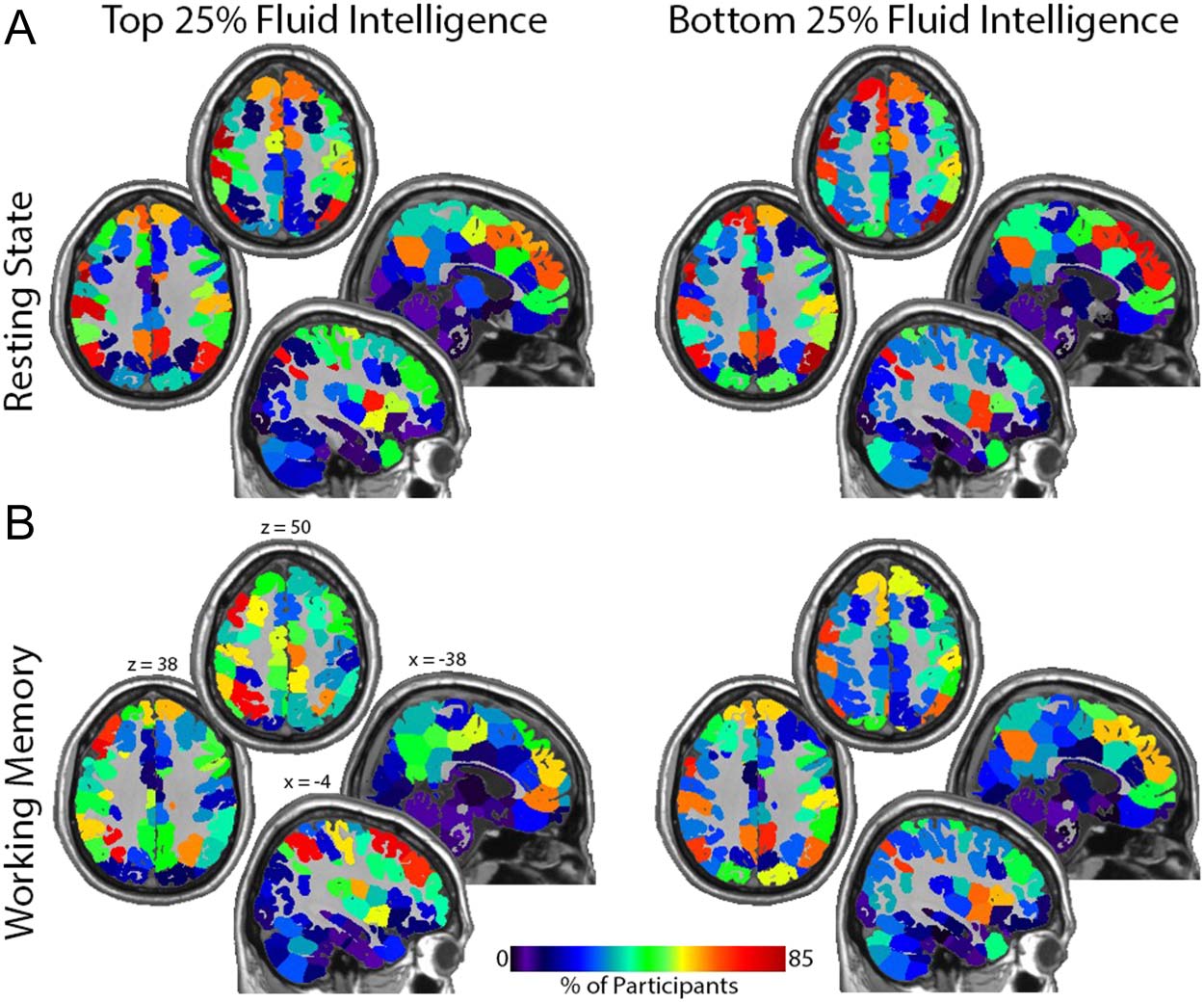
Figure 7. Maps showing the location of network hubs (top 20% degree) for the top and bottom
intelligence quartiles. (A) Resting state: note the high incidence of hubs in the medial and lateral
parietal cortex and the medial frontal lobe in both groups. These regions are central components of
the default mode network. The significant association between degree location and intelligence is
demonstrated by the reduction in default mode network hubs with increases in fluid intelligence. (B)
Working memory: hubs are more concentrated in the central executive network as intelligence
increases. This shift is best appreciated in the top quartile group where hubs are concentrated in
lateral rather than medial frontal regions and in more superior parietal areas. Note that the hubs
remain fairly localized to the default mode network for the bottom quartile group. Each quadrant
shows two axial and two sagittal images. The Montreal Neurological Institute (MNI) coordinates
shown in the bottom left quadrant apply to all quadrants. Calibration bar applies to all images.
prominent in the areas of the default mode network even during the working memory task,
although the magnitude of the overlap is lower. The CEN maps onto what is sometimes
referred to as the fronto-parietal network. There is evidence that the density of structural con-
nectivity in that network predicts working memory capacity, with higher capacity individuals
exhibiting greater connectivity (Ekman et al., 2016).
Among the confounding variables, some had significant relationships with all distance met-
rics except for the JD (age and alcohol abuse within resting state). There were also certain
variables that were significant for one fMRI task but not the other (alcohol abuse, BMI, and
education). Some covariates were significant for all location-specific metrics but not for the
distance between distribution comparator KS statistic (BMI – rest; gender – rest and working
memory; age, fluid intelligence and race – working memory). This was indicative of a detect-
able difference in location-specific degree, but not in distribution.
In addition to our primary analysis detailed above, we also examined these relationships
with respect to differences in modularity between individuals employing scaled inclusivity,
given that modularity analyses can capture the spatial distribution of intrinsic brain networks
that are associated with various cognitive tasks (Moussa et al., 2012). The description of this
secondary analysis, along with the corresponding results and figures can be found in the Sup-
porting Information.
Network Neuroscience
63
A regression framework for brain network distance metrics
DISCUSSION
It is of great interest to compare differences in fMRI connection matrices between individuals
by covariates. Our previous work developed a novel permutation testing framework to detect
differences between two groups (Simpson et al., 2013b). Here, we advanced that work to
allow both assessing differences by continuous phenotypes and controlling for confounding
variables. We proposed an innovative regression framework to relate distances between brain
network features to functions of absolute differences in continuous covariates and indicators of
difference for categorical variables, and explored several similarity metrics for comparing dis-
tances between connection matrices. The KS statistic measures how different distributions of
topological properties vary between two individuals. Key node metrics (like the JD) quantify
how much the spatial location of key brain network regions differs between two networks. The
Euclidean norm (along with Canberra, Minkowski, Weighted Jaccard, etc.) measures whether
the spatial location of degree-weighted brain network regions differ. Several additional
similarity/dissimilarity metrics mentioned below were also assessed, but details were left out
for the sake of brevity. The binary network-based metrics left out included the Similarity
Matching, Russel-Rao, Kulczynski, and Baroni-Urbani and Buser. Little difference or improve-
ment was found (in Simulations 1–3) for any of these metrics when compared with the
commonly used Jaccard. The weighted network-based metrics left out included the Weighted
Jaccard, Manhattan, and Maximum distances, with none different or better than the already
presented Canberra or Minkowski distances (when tested in Simulations 1–3). Any other dis-
tance or similarity metrics could be used. Future work might include (a) testing other metrics,
(b) further investigating the behavior of the Jaccard metric, and (c) taking a deeper dive into
understanding when and how to choose a distance metric.
Several standard methods for estimation and inference were adapted to fit into our regres-
sion framework: standard F test, F test with ILE, GLS, and permutation (inference only). All
combinations of these approaches and the distance metrics were assessed via four simulation
scenarios. The KS statistic was found to have low power (relative to the other distance metrics)
when testing location-specific differences. However, if interested in comparing distributions,
the KS statistic was preferred. The JD did not have consistent or predictable power, and further
work should investigate the reasons underlying this. An argument could be made for several
different distance metrics when detecting location-specific differences between degree vec-
tors, as they had very similar results. We prefer to use Euclidean distance, as it is commonly
recognized and had the best overall (slightly) type I error control and power. Future work could
include a further investigation into how and when to choose specific distance metrics. Regard-
ing the comparison of estimation and testing methods (standard F test, F test with ILE, GLS,
permutation), we recommend the F test with ILE as it was among the best at controlling type
I error and providing sufficient power while also being computationally inexpensive.
An FGLS approach for estimation and inference was proposed in this framework.
Although it is slower (computationally) than the standard F test with ILE, the FGLS approach
performed just as well as the recommended Standard F test with ILE (in terms of type I and
type II error).
An analysis of the HCP data was completed using the standard F test with ILE and several
distance metrics. After adjusting for the other confounding variables, the covariate of interest,
fluid intelligence, was significantly related with all distance metrics (KS, Jaccard, Euclidean) for
both fMRI tasks (resting state, working memory). This analysis suggested the hubs become
more strongly concentrated in default mode network regions at rest with decreasing fluid
intelligence. As intelligence increases, there are greater shifts in the spatial locations of hubs
Network Neuroscience
64
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
t
/
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
from the default mode network at rest to the central executive network during a working
memory task.
Many existing methods exist for relating network metrics and phenotypes. Such methods
include, but are not limited to traditional network models (e.g., exponential random graph
models; Lehmann et al., 2021; Simpson et al., 2011, 2012), tensor regression works on brain
network (e.g., Zhang et al., 2018, 2019), Bayesian approaches (e.g., Dai & Guo, 2017; Wang
et al., 2017), statistical learning techniques (Craddock et al., 2015; Varoquaux & Craddock,
2013; Xia et al., 2020), and testing based on distance correlation (Székely et al., 2007; Székely
& Rizzo, 2009). We believe our method most closely relates to MDMR. MDMR tests the sig-
nificance of associations of response profile (dis)similarities and a set of predictors. Originally
this was done using only permutation (Anderson, 2001) but has been extended to analytic
p values as well (McArtor et al., 2017). Via simulation, a comparison to our regression frame-
work showed that MDMR performs relatively well for the Euclidean and Jaccard distances (as
well as other Minkowski distances—see Supporting Information) in terms of power. However,
its type I error rate properties were not well understood. For many covariates, close to 0% of
tests had p values less than 0.05. Further investigation should be done to better understand this
property. Furthermore, MDMR was not able to detect differences on either the KS statistic or JI.
Our framework vastly outperformed MDMR for these two metrics for all simulation scenarios
except for Simulation 4/Jaccard in which the results were comparable (see section Jaccard
distance and Jaccard index in Results for an explanation).
We have developed a testing framework that detects whether the spatial location of key
brain network regions and distributions of topological properties differ by phenotype (contin-
uous and discrete) after controlling for confounding variables in static networks. More gener-
ally, this framework allows relating distances between networks (e.g., Jaccard, KS distance) to
covariates of interest. Our chosen method, F test with ILE, is computationally inexpensive,
generally interpretable, and well understood by most scientists. We believe this adds a con-
venient tool to the neuroscience toolbox. Future work plans to extend this approach by cre-
ating a dynamic network analog that uncovers whether within- and across-task time-varying
changes in these spatial and distributional patterns differ by phenotype.
ACKNOWLEDGMENTS
The authors thank Hongtu Zhu, Professor of Biostatistics at University of North Carolina at
Chapel Hill, for suggesting that we assess the logarithmic transformations of the distance met-
rics. We also thank Dale Dagenbach, Professor of Psychology at Wake Forest University, for
his insights into interpreting our HCP results regarding IQ.
SUPPORTING INFORMATION
Supporting information for this article is available at https://doi.org/10.1162/netn_a_00214.
Simulation and HCP code is available: https://github.com/applebrownbetty/braindist_regression.
HCP data is publicly available for download.
AUTHOR CONTRIBUTIONS
Chalmer E. Tomlinson: Conceptualization; Data curation; Formal analysis; Investigation;
Methodology; Project administration; Resources; Supervision; Validation; Visualization;
Writing – original draft; Writing – review & editing. Paul J. Laurienti: Data curation; Resources;
Visualization; Writing – original draft; Writing – review & editing. Robert G. Lyday: Data
curation; Visualization; Writing – original draft; Writing – review & editing. Sean L. Simpson:
Network Neuroscience
65
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
t
/
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
Conceptualization; Data curation; Formal analysis; Funding acquisition; Investigation;
Methodology; Project administration; Resources; Supervision; Validation; Visualization;
Writing – original draft; Writing – review & editing.
FUNDING INFORMATION
Sean L. Simpson, National Institute of Biomedical Imaging and Bioengineering (https://dx.doi
.org/10.13039/100000070), Award ID: R01EB024559. Sean L. Simpson, Wake Forest Clinical
and Translational Science Institute, Award ID: NCATS UL1TR001420.
REFERENCES
Aitken, A. C. (1936). On least-squares and linear combinations of
observations. Proceedings of the Royal Society of Edinburgh, 55,
42–48. https://doi.org/10.1017/S0370164600014346
Anderson, M. J. (2001). A new method for non-parametric multivar-
iate analysis of variance. Austral Ecology, 26, 32–46. https://doi
.org/10.1111/J.1442-9993.2001.01070.PP.X
Bassett, D. S., & Bullmore, E. T. (2009). Human brain networks in
health and disease. Current Opinion in Neurology. https://doi.org
/10.1097/ WCO.0b013e32832d93dd, PubMed: 19494774
Bates, D., Mächler, M., Bolker, B., & Walker, S. (2015). Fitting linear
mixed-effects models using lme4. Journal of Statistical Software,
67, 1–48. https://doi.org/10.18637/JSS.V067.I01
Burdette, J. H., Laurienti, P. J., Espeland, M. A., Morgan, A., Telesford,
Q., Vechlekar, C. D., Hayasaka, S., Jennings, J. M., Katula, J. A.,
Kraft, R. A., & Rejeski, W. J. (2010). Using network science to
evaluate exercise-associated brain changes in older adults.
Frontiers in Aging Neuroscience, 2. https://doi.org/10.3389
/fnagi.2010.00023, PubMed: 20589103
Craddock, R. C., Tungaraza, R. L., & Milham, M. P. (2015).
Connectomics and new approaches for analyzing human brain
functional connectivity. Gigascience, 4. https://doi.org/10.1186
/S13742-015-0045-X, PubMed: 25810900
Dai, T., & Guo, Y. (2017). Predicting individual brain functional
connectivity using a Bayesian hierarchical model. NeuroImage,
147, 772–787. https://doi.org/10.1016/j.neuroimage.2016.11
.048, PubMed: 27915121
Deuker, L., Bullmore, E. T., Smith, M., Christensen, S., Nathan, P. J.,
Rockstroh, B., & Bassett, D. S. (2009). Reproducibility of graph
metrics of human brain functional networks. NeuroImage, 47,
1460–1468. https://doi.org/10.1016/j.neuroimage.2009.05.035,
PubMed: 19463959
Ekman, M., Fiebach, C. J., Melzer, C., Tittgemeyer, M., & Derrfuss,
J. (2016). Different roles of direct and indirect frontoparietal
pathways for individual working memory capacity. Journal of
Neuroscience, 36, 2894–2903. https://doi.org/10.1523
/JNEUROSCI.1376-14.2016, PubMed: 26961945
Fornito, A., Zalesky, A., Pantelis, C., & Bullmore, E. T. (2012).
Schizophrenia, neuroimaging and connectomics. NeuroImage.
https://doi.org/10.1016/j.neuroimage.2011.12.090, PubMed:
22387165
Fraiman, D., Balenzuela, P., Foss, J., & Chialvo, D. R. (2009). Ising-
like dynamics in large-scale functional brain networks. Physical
Review E, 79, 061922. https://doi.org/10.1103/ PhysRevE.79
.061922, PubMed: 19658539
Freedman, D., & Lane, D. (1983). A nonstochastic interpretation
of reported significance levels. Journal of Business & Economic
Statistics, 1, 292–298. https://doi.org/10.1080/07350015.1983
.10509354
Frossard, J., & Renaud, O. (2019a). permuco: Permutation tests for
regression, (repeated measures) ANOVA/ANCOVA and compar-
ison of signals [Computer software manual].
Frossard, J., & Renaud, O. (2019b). Permutation tests for regression,
ANOVA and comparison of signals: The permuco package. R
Packag. Version 1.1.0.
Glasser, M. F., Sotiropoulos, S. N., Wilson, J. A., Coalson, T. S.,
Fischl, B., Andersson, J. L., Xu, J., Jbabdi, S., Webster, M.,
Polimeni, J. R., Van Essen, D. C., & Jenkinson, M. (2013). The
minimal preprocessing pipelines for the Human Connectome
Project. NeuroImage, 80, 105–124. https://doi.org/10.1016/j
.neuroimage.2013.04.127, PubMed: 23668970
Hagmann, P., Cammoun, L., Gigandet, X., Meuli, R., Honey, C. J.,
Van Wedeen, J., & Sporns, O. (2008). Mapping the structural core
of human cerebral cortex. PLoS Biology, 6, 1479–1493. https://
doi.org/10.1371/journal.pbio.0060159, PubMed: 18597554
Hayasaka, S., & Laurienti, P. J. (2010). Comparison of characteris-
tics between region-and voxel-based network analyses in
resting-state fMRI data. NeuroImage, 50, 499–508. https://doi
.org/10.1016/j.neuroimage.2009.12.051, PubMed: 20026219
Joyce, K. E., Laurienti, P. J., Burdette, J. H., & Hayasaka, S. (2010). A
new measure of centrality for brain networks. PLoS One, 5. https://
doi.org/10.1371/journal.pone.0012200, PubMed: 20808943
Kherad-Pajouh, S., & Renaud, O. (2010). An exact permutation
method for testing any effect in balanced and unbalanced fixed
effect ANOVA. Computational Statistics & Data Analysis, 54,
1881–1893. https://doi.org/10.1016/j.csda.2010.02.015
Kherad-Pajouh, S., & Renaud, O. (2014). A general permutation
approach for analyzing repeated measures ANOVA and mixed-
model designs. Statistical Papers, 56, 947–967. https://doi.org/10
.1007/s00362-014-0617-3
Kolmogorov, A. (1933). Sulla determinazione empirica di una lgge di
distribuzione. Giornale dellʼIstituto Italiano degli Attuari, 4, 83–91.
Lance, G. N., & Williams, W. T. (1966). Computer programs for
hierarchical polythetic classification (“similarity analyses”).
Computer Journal, 9, 60–64. https://doi.org/10.1093/comjnl/9.1.60
Network Neuroscience
66
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
/
t
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
Lehmann, B. C. L., Henson, R. N., Geerligs, L., Cam-CAN, &
White, S. R. (2021). Characterising group-level brain connec-
tivity: A framework using Bayesian exponential random graph
models. NeuroImage, 225. https://doi.org/10.1016/j.neuroimage
.2020.117480, PubMed: 33099009
Liu, Y., Liang, M., Zhou, Y., He, Y., Hao, Y., Song, M., Yu, C., Liu,
H., Liu, Z., & Jiang, T. (2008). Disrupted small-world networks in
schizophrenia. Brain, 131, 945–961. https://doi.org/10.1093
/brain/awn018, PubMed: 18299296
McArtor, D. B. (2018). MDMR: Multivariate Distance Matrix
Regression. R package version 0.5.1. https://CRAN.R-project
.org/package=MDMR [cran.r-project.org].
McArtor, D. B., Lubke, G. H., & Bergeman, C. S. (2017). Extending
multivariate distance matrix regression with an effect size mea-
sure and the asymptotic null distribution of the test statistic. Psy-
chometrika, 82, 1052. https://doi.org/10.1007/S11336-016-9527
-8, PubMed: 27738957
Meunier, D., Achard, S., Morcom, A., & Bullmore, E. (2009a). Age-
related changes in modular organization of human brain func-
tional networks. NeuroImage, 44, 715–723. https://doi.org/10
.1016/j.neuroimage.2008.09.062, PubMed: 19027073
Meunier, D., Lambiotte, R., Fornito, A., Ersche, K. D., & Bullmore,
E. T. (2009b). Hierarchical modularity in human brain functional
networks. Frontiers in Neuroinformatics, 3. https://doi.org/10
.3389/neuro.11.037.2009, PubMed: 19949480
Moussa, M. N., Steen, M. R., Laurienti, P. J., & Hayasaka, S. (2012).
Consistency of network modules in resting-state fMRI connec-
tome data. PLoS One, 7. https://doi.org/10.1371/journal.pone
.0044428, PubMed: 22952978
Moussa, M. N., Vechlekar, C. D., Burdette, J. H., Steen, M. R.,
Hugenschmidt, C. E., & Laurienti, P. J. (2011). Changes in cogni-
tive state alter human functional brain networks. Frontiers in
Human Neuroscience. https://doi.org/10.3389/fnhum.2011
.00083, PubMed: 21991252
Parente, F., Frascarelli, M., Mirigliani, A., Di Fabio, F., Biondi, M., &
Colosimo, A. (2017). Negative functional brain networks. Brain
Imaging and Behavior, 12, 467–476. https://doi.org/10.1007
/S11682-017-9715-X, PubMed: 28353136
Pruim, R. H. R., Mennes, M., Van Rooij, D., Llera, A., Buitelaar,
J. K., & Beckmann, C. F. (2015). ICA-AROMA: A robust
ICA-based strategy for removing motion artifacts from fMRI data.
NeuroImage, 112, 267–277. https://doi.org/10.1016/j
.neuroimage.2015.02.064, PubMed: 25770991
Rombouts, S. A. R. B, Barkhof, F., Goekoop, R., Stam, C. J., &
Scheltens, P. (2005). Altered resting state networks in mild cogni-
tive impairment and mild Alzheimer’s disease: An fMRI study.
Human Brain Mapping, 26, 231–239. https://doi.org/10.1002
/hbm.20160, PubMed: 15954139
Rzucidlo, J. K., Roseman, P. L., Laurienti, P. J., & Dagenbach, D. (2013).
Stability of whole brain and regional network topology within
and between resting and cognitive states. PLoS One, 8. https://
doi.org/10.1371/journal.pone.0070275, PubMed: 23940554
Saberi, M., Khosrowabadi, R., Khatibi, A., Misic, B., & Jafari, G.
(2021). Topological impact of negative links on the stability of
resting-state brain network. Scientific Reports, 11. https://doi.org
/10.1038/S41598-021-81767-7, PubMed: 33500525
Schwarz, A. J., & McGonigle, J. (2011). Negative edges and soft
thresholding in complex network analysis of resting state
functional connectivity data. NeuroImage, 55, 1132–1146. https://
doi.org/10.1016/j.neuroimage.2010.12.047, PubMed: 21194570
Shen, X., Tokoglu, F., Papademetris, X., & Constable, R. T. (2013).
Groupwise whole-brain parcellation from resting-state fMRI data
for network node identification. NeuroImage, 82, 403–415.
https://doi.org/10.1016/j.neuroimage.2013.05.081, PubMed:
23747961
Simpson, S. L., Bowman, F. D. B., & Laurienti, P. J. (2013a). Ana-
lyzing complex functional brain networks: Fusing statistics and
network science to understand the brain. Statistical Surveys, 7,
1–36. https://doi.org/10.1214/13-SS103, PubMed: 25309643
Simpson, S. L., Hayasaka, S., Laurienti, P. J. (2011). Exponential ran-
dom graph modeling for complex brain networks. PLoS One, 6.
https://doi.org/10.1371/journal.pone.0020039, PubMed:
21647450
Simpson, S. L., Lyday, R. G., Hayasaka, S., Marsh, A. P., & Laurienti,
P. J. (2013b). A permutation testing framework to compare groups
of brain networks. Frontiers in Computational Neuroscience, 7,
1–13. https://doi.org/10.3389/fncom.2013.00171, PubMed:
24324431
Simpson, S. L., Moussa, M. N., & Laurienti, P. J. (2012). An expo-
nential random graph modeling approach to creating
group-based representative whole-brain connectivity networks.
NeuroImage, 60, 1117–1126. https://doi.org/10.1016/j
.neuroimage.2012.01.071, PubMed: 22281670
Smirnov, N. (1948). Table for estimating the goodness of fit of
empirical distributions. Annals of Mathematical Statistics, 19,
279–281. https://doi.org/10.1214/aoms/1177730256
Stam, C. J., Jones, B. F., Nolte, G., Breakspear, M., & Scheltens, P.
(2007). Small-world networks and functional connectivity in
Alzheimer’s disease. Cerebral Cortex, 17, 92–99. https://doi.org
/10.1093/cercor/bhj127, PubMed: 16452642
Székely, G. J., & Rizzo, M. L. (2009). Brownian distance covari-
ance. Annals of Applied Statistics, 3, 1236–1265. https://doi.org
/10.1214/09-AOAS312, PubMed: 20574547
Székely, G. J., Rizzo, M. L., & Bakirov, N. K. (2007). Measuring and
testing dependence by correlation of distances. Annals of Statistics,
35, 2769–2794. https://doi.org/10.1214/009053607000000505
van den Heuvel, M. P., Stam, C. J., Boersma, M., & Hulshoff Pol,
H. E. (2008). Small-world and scale-free organization of
voxel-based resting-state functional connectivity in the human
brain. NeuroImage, 43, 528–539. https://doi.org/10.1016/j
.neuroimage.2008.08.010, PubMed: 18786642
Van Essen, D. C., Smith, S. M., Barch, D. M., Behrens, T. E. J.,
Yacoub, E., & Ugurbil, K. (2013). The WU-Minn Human Connec-
tome Project: An overview. NeuroImage, 80, 62–79. https://doi
.org/10.1016/j.neuroimage.2013.05.041, PubMed: 23684880
Van Essen, D. C., Ugurbil, K., Auerbach, E., Barch, D., Behrens,
T. E. J., Bucholz, R., Chang, A., Chen, L., Corbetta, M., Curtiss,
S. W., Della Penna, S., Feinberg, D., Glasser, M. F., Harel, N.,
Heath, A. C., Larson-Prior, L., Marcus, D., Michalareas, G.,
Moeller, S., Oostenveld, R., Petersen, S. E., Prior, F., Schlaggar,
B. L., Smith, S. M., Snyder, A. Z., Xu, J., & Yacoub, E. (2012). The
Human Connectome Project: A data acquisition perspective.
NeuroImage. https://doi.org/10.1016/j.neuroimage.2012.02.018,
PubMed: 22366334
Varoquaux, G., & Craddock, R. C. (2013). Learning and comparing
functional connectomes across subjects. NeuroImage, 80, 405–415.
Network Neuroscience
67
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
/
/
t
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
.
t
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
A regression framework for brain network distance metrics
https://doi.org/10.1016/J.NEUROIMAGE.2013.04.007, PubMed:
23583357
Wang, L., Durante, D., Jung, R. E., & Dunson, D. B. (2017). Bayesian
network–response regression. Bioinformatics, 33, 1859–1866.
https://doi.org/10.1093/ BIOINFORMATICS/ BTX050, PubMed:
28165112
Xia, C. H., Ma, Z., Cui, Z., Bzdok, D., Thirion, B., Bassett, D. S.,
Satterthwaite, T. D., Shinohara, R. T., & Witten, D. M. (2020).
Multi-scale network regression for brain-phenotype associations.
Human Brain Mapping, 41, 2553–2566. https://doi.org/10.1002
/hbm.24982, PubMed: 32216125
Yuan, K., Qin, W., Liu, J., Guo, Q., Dong, M., Sun, J., Zhang, Y.,
Liu, P., Wang, W., Wang, Y., Li, Q., Yang, W., von Deneen,
K. M., Gold, M. S., Liu, Y., & Tian,
(2010). Altered
small-world brain functional networks and duration of heroin
use in male abstinent heroin-dependent individuals. Neuroscience
Letters, 477, 37–42. https://doi.org/10.1016/j.neulet.2010.04.032,
PubMed: 20417253
J.
Zhang, J., Sun, W. W., & Li, L. (2018). Network response regression
for modeling population of networks with covariates. arXiv
preprint arXiv:1810.03192v2.
Zhang, Z., Allen, G. I., Zhu, H., & Dunson, D. (2019). Tensor
network factorizations: Relationships between brain structural
connectomes and traits. NeuroImage, 197, 330–343. https://
doi.org/10.1016/j.neuroimage.2019.04.027, PubMed:
31029870
l
D
o
w
n
o
a
d
e
d
f
r
o
m
h
t
t
p
:
/
/
d
i
r
e
c
t
.
m
i
t
.
t
/
/
e
d
u
n
e
n
a
r
t
i
c
e
–
p
d
l
f
/
/
/
/
/
6
1
4
9
1
9
8
4
2
1
8
n
e
n
_
a
_
0
0
2
1
4
p
d
t
.
f
b
y
g
u
e
s
t
t
o
n
0
7
S
e
p
e
m
b
e
r
2
0
2
3
Network Neuroscience
68